Learn More
general transcription factor IIH, polypeptide 2, 44kDa, Mouse, Polyclonal Antibody, Abnova™
Shop All Abnova Corporation ProductsDescription
This gene is part of a 500kb inverted duplication on chromosome 5q13. This duplicated region contains at least four genes and repetitive elements which make it prone to rearrangements and deletions. The repetitiveness and complexity of the sequence have also caused difficulty in determining the organization of this genomic region. This gene is within the telomeric copy of the duplication. Deletion of this gene sometimes accompanies deletion of the neighboring SMN1 gene in spinal muscular atrophy (SMA) patients but it is unclear if deletion of this gene contributes to the SMA phenotype. This gene encodes the 44kDa subunit of RNA polymerase II transcription initiation factor IIH which is involved in basal transcription and nucleotide excision repair. Transcript variants for this gene have been described, but their full length nature has not been determined. A second copy of this gene within the centromeric copy of the duplication has been described in the literature. It is reported to be different by either two or four base pairs; however, no sequence data is currently available for the centromeric copy of the gene. [provided by RefSeq
Specifications
Specifications
| Antigen | general transcription factor IIH, polypeptide 2, 44kDa |
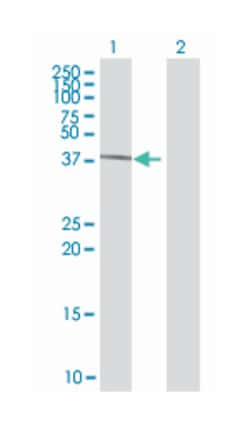
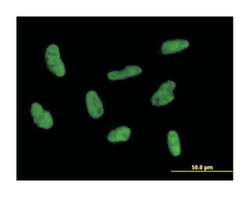
| Applications | Immunofluorescence, Western Blot |
| Classification | Polyclonal |
| Conjugate | Unconjugated |
| Description | Mouse polyclonal antibody raised against a full-length human GTF2H2 protein. |
| Formulation | No additive |
| Gene | GTF2H2 |
| Gene Accession No. | NM_001515.2 |
| Gene Alias | BTF2/BTF2P44/MGC102806/T-BTF2P44/TFIIH |
| Gene Symbols | GTF2H2 |
| Show More |
By clicking Submit, you acknowledge that you may be contacted by Fisher Scientific in regards to the feedback you have provided in this form. We will not share your information for any other purposes. All contact information provided shall also be maintained in accordance with our Privacy Policy.