Learn More
Invitrogen™ MST2 Monoclonal Antibody (1E9)


Description
Peptide Sequence: MEQPPAPKSK LKKLSEDSLT KQPEEVFDVL EKLGEGSYGS VFKAIHKESG QVVAIKQVPV ESDLQEIIKE ISIMQQCDSP YVVKYYGSYF KNTDLWIVME YCGAGSVSDI IRLRNKTLIE DEIATILKST LKGLEYLHFM RKIHRDIKAG NILLNTEGHA KLADFGVAGQ LTDTMAKRNT VIGTPFWMAP EVIQEIGYNC VADIWSLGIT SIEMAEGKPP YADIHPMRAI FMIPTNPPPT FRKPELWSDD FTDFVKKCLV KNPEQRATAT QLLQHPFIKN AKPVSILRDL ITEAMEIKAK RHEEQQRELE EEEENSDEDE LDSHTMVKTS VESVGTMRAT STMSEGAQTM IEHNSTMLES DLGTMVINSE DEEEEDGTMK RNATSPQVQR PSFMDYFDKQ DFKNKSHENC NQNMHEPFPM SKNVFPDNWK VPQDGDFDFL KNLSLEELQM RLKALDPMME REIEELRQRY TAKRQPILDA MDAKKRRQQN F.

Specifications
Specifications
| Antigen | MST2 |
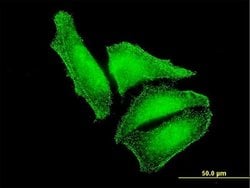
| Applications | ELISA, Immunocytochemistry |
| Classification | Monoclonal |
| Clone | 1E9 |
| Concentration | 0.2 mg/mL |
| Conjugate | Unconjugated |
| Formulation | PBS with no preservative; pH 7.4 |
| Gene | STK3 |
| Gene Accession No. | Q13188 |
| Gene Alias | 0610042I06Rik; KB-1458E12.1; KRS1; Mammalian STE20-like protein kinase 2; mess1; MST; Mst2; MST-2; MST2/C; MST2/N; Mst3; protein kinase homolog; serine/threonine kinase 3; serine/threonine kinase 3 (STE20 homolog, yeast); serine/threonine kinase 3 (Ste20, yeast homolog); serine/threonine-protein kinase 3; Serine/threonine-protein kinase 3 20kDa subunit; Serine/threonine-protein kinase 3 36kDa subunit; Serine/threonine-protein kinase Krs-1; Ste20; STE20-like kinase MST2; STK3; wu:fc19e11; zgc:55383 |
| Show More |
By clicking Submit, you acknowledge that you may be contacted by Fisher Scientific in regards to the feedback you have provided in this form. We will not share your information for any other purposes. All contact information provided shall also be maintained in accordance with our Privacy Policy.