Learn More
Description
The protein encoded by this gene is a member of the fibroblast growth factor receptor family, where amino acid sequence is highly conserved between members and throughout evolution. FGFR family members differ from one another in their ligand affinities and tissue distribution. A full-length representative protein consists of an extracellular region, composed of three immunoglobulin-like domains, a single hydrophobic membrane-spanning segment and a cytoplasmic tyrosine kinase domain. The extracellular portion of the protein interacts with fibroblast growth factors, setting in motion a cascade of downstream signals, ultimately influencing mitogenesis and differentiation. This particular family member is a high-affinity receptor for acidic, basic and/or keratinocyte growth factor, depending on the isoform. Mutations in this gene are associated with Crouzon syndrome, Pfeiffer syndrome, Craniosynostosis, Apert syndrome, Jackson-Weiss syndrome, Beare-Stevenson cutis gyrata syndrome, Saethre-Chotzen syndrome, and syndromic craniosynostosis. Multiple alternatively spliced transcript variants encoding different isoforms have been noted for this gene. [provided by RefSeq
Specifications
Specifications
| Antigen | FGFR2 |
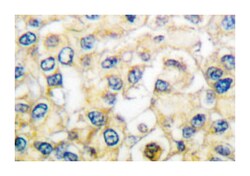
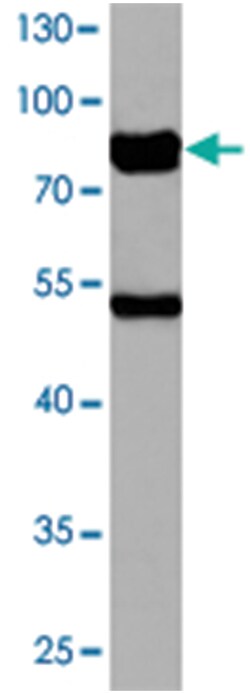
| Applications | Immunohistochemistry (PFA fixed), Western Blot |
| Classification | Polyclonal |
| Conjugate | Unconjugated |
| Description | Rabbit polyclonal antibody raised against synthetic peptide of FGFR2. |
| Dilution | Western Blot (1:500-1:1000) Immunohistochemistry (1:50-1:200) The optimal working dilution should be determined by the end user. |
| Formulation | In PBS, pH 7.2 (0.09% sodium azide) |
| Gene | FGFR2 |
| Gene Accession No. | P21802 |
| Gene Alias | BEK/BFR-1/CD332/CEK3/CFD1/ECT1/FLJ98662/JWS/K-SAM/KGFR/TK14/TK25 |
| Show More |
For Research Use Only
By clicking Submit, you acknowledge that you may be contacted by Fisher Scientific in regards to the feedback you have provided in this form. We will not share your information for any other purposes. All contact information provided shall also be maintained in accordance with our Privacy Policy.