Learn More
Description
Biological processes such as transmembrane signaling and receptor mediated endocytosis revolve around the function of cell surface receptors. A network of molecular machinery directs the intracellular trafficking of receptors during their biosynthesis and mediates signaling downstream of receptors. The sorting nexins (SNX1, SNX1A, SNX2, SNX3, and SNX4) are a family of intracellular proteins that are thought to direct the sorting of receptor proteins. The SNX proteins contain a conserved 100 amino acid region termed the phox homology (PX)
domain and are part of a family of hydrophilic proteins which includes S. cerevisiae proteins that function in protein sorting. SNX1, SNX2 and SNX4 associate predominantly with membranes and bind transmembrane receptors such as those for EGF, PDGF, and insulin. SNX1 directs the EGF receptor to the lysosomes for degradation. SNX2 forms homomeric complexes and heteromeric complexes with SNX1, SNX1A, and SNX4. These complexes are thought to be necessary for efficient protein sorting. Thus, SNX1 and other sorting nexins are thought to play important roles in the specificity of protein trafficking to and from the plasma membrane.
Host Species: Mouse
Clone: 51
Isotype: IgG1
Species Reactivity: Human
Immunogen: Human SNX1 aa. 1-108
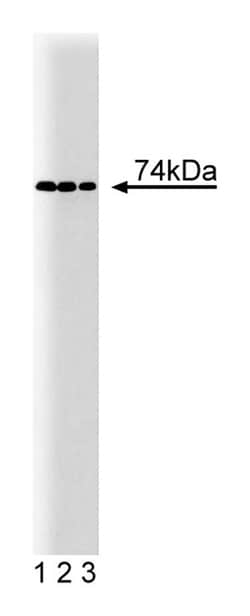
Formula Weight [Chemical]: 74kDa
Immunofluorescence, Western Blotting

Specifications
Specifications
| Antigen | SNX1 |
| Applications | Western Blot |
| Classification | Monoclonal |
| Clone | 51 |
| Concentration | 250μg/mL |
| Conjugate | Unconjugated |
| Formulation | Aqueous buffered solution containing BSA, glycerol, and ≤0.09% sodium azide. |
| Host Species | Mouse |
| Immunogen | Human SNX1 aa. 1-108 |
| Purification Method | Affinity Purified |
| Show More |
Safety and Handling
By clicking Submit, you acknowledge that you may be contacted by Fisher Scientific in regards to the feedback you have provided in this form. We will not share your information for any other purposes. All contact information provided shall also be maintained in accordance with our Privacy Policy.