Promotional price valid on web orders only. Your contract pricing may differ. Interested in signing up for a dedicated account number?
Learn More
Learn More
BTF3L4 Rabbit anti-Human, Mouse, Rat, Polyclonal, Proteintech


Rabbit Polyclonal Antibody
Supplier: Proteintech Group Inc 165001AP150UL

Description
The initiation of gene transcription involves the ordered assembly of a multiprotein complex on proximal promoter elements such as the TATA box. In addition to RNA polymerase II (Pol II), the transcription factor class II (TFII) family of proteins are required for initiation of transcription, as the first step in the formation of this initiation complex is the stable binding of TFIID to the TATA box. An additional TFII-related protein, BTF3, does not directly associate with the proximal promoter, but rather forms a stable complex with Pol II and facilitates Pol II assembly into the complex. The BTF3 gene is ubiquitously expressed and encodes two protein isoforms, BTF3a and BTF3b, which are produced from alternative splicing. The BTF3 proteins are identical, except that BTF3b lacks the first 44 amino acids at the N-terminal of BTF3a. As a consequence of this deletion, BTF3b is unable to induce transcription, despite being able to bind Pol II. Additionally, BTF3a and BTF3b associate with the widely expressed protein kinase casein kinase II. Casein kinase II phosphorylates BTF3a, as well as TFIIB, and is required for the efficient transcription of the tRNA and 55 rRNA genes by Pol III. BTF3 belongs to the NAC-beta family, which includes several related proteins, such as BTF3L1, BTF3L2, BTF3L3 and BTF3L4.Specifications
| BTF3L4 | |
| Polyclonal | |
| Unconjugated | |
| BTF3L4 | |
| BTF3L4, RP4 800M22.5 | |
| Rabbit | |
| Antigen Affinity Chromatography | |
| RUO | |
| 366434, 70533, 91408 | |
| -20°C | |
| Liquid |
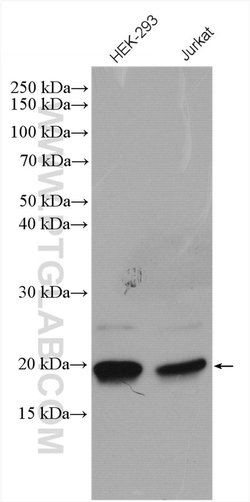
| Western Blot | |
| 0.2 mg/mL | |
| PBS with 50% glycerol and 0.1% sodium azide; pH 7.3 | |
| Q96K17, Q9CQH7 | |
| Btf3l4 | |
| BTF3L4 Fusion Protein Ag9666 | |
| 150 μL | |
| Primary | |
| Human, Mouse, Rat | |
| Antibody | |
| IgG |
Product Content Correction
The Fisher Scientific Encompass Program offers items which are not part of our distribution portfolio. These products typically do not have pictures or detailed descriptions. However, we are committed to improving your shopping experience. Please use the form below to provide feedback related to the content on this product.
Product Title
Spot an opportunity for improvement?Share a Content Correction