Learn More
Invitrogen™ Phospho-STAT4 (Tyr693) Monoclonal Antibody (4LURPIE), PE, eBioscience™, Invitrogen™

Description
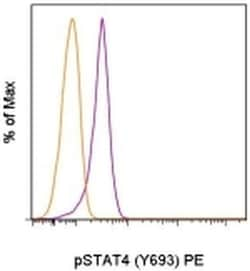
Description: This 4LURPIE monoclonal antibody recognizes human and mouse signal transducer and activator of transcription 4 (STAT4) when phosphorylated on tyrosine 693 (Y693). STAT proteins are activated by ligand binding to cytokine receptors that associate with Janus kinase (JAK) family members. Following their phosphorylation by JAK proteins, STAT proteins translocate to the nucleus where they bind to DNA and regulate transcription of specific genes in a cell type- and cytokine-specific manner. STAT4 is activated by IL-12 or type 1 IFN (IFN alpha, IFN beta). STAT4 is required for optimal Th1 differentiation in vivo. Interestingly, although IFN alpha can lead to phosphorylation of STAT4 in mouse T cells, the phosphorylation is much weaker than that induced by IL-12 and is transient. Thus, IFN alpha phosphorylation of STAT4 does not contribute to STAT4-mediated Th1 development or IFN gamma production by Th1 cells in mice. In contrast, IFN alpha does promote Th1 development and function of human CD4+ T cells. Specificity of this 4LURPIE clone was determined by ELISA and flow cytometry. Applications Reported: This 4LURPIE antibody has been reported for use in intracellular staining followed by flow cytometric analysis. Applications Tested: This 4LURPIE antibody has been pre-titrated and tested by intracellular staining followed by flow cytometric analysis of stimulated normal human peripheral blood cells. This can be used at 5 μL (0.25 μg) per test.

Specifications
Specifications
| Antigen | Phospho-STAT4 (Tyr693) |
| Applications | Flow Cytometry |
| Classification | Monoclonal |
| Clone | 4LURPIE |
| Concentration | 5 μL/Test |
| Conjugate | PE |
| Formulation | PBS with BSA and 0.09% sodium azide; pH 7.2 |
| Gene | STAT4 |
| Gene Accession No. | P42228, Q14765 |
| Gene Alias | CIS4; HSPC060; OTTHUMP00000163553; Signal transducer and activator of transcription 4; SLEB11; SOCS4; SOCS6; SSI4; STAI4; STAT; Stat4; STATI4 |
| Show More |
For Research Use Only.
By clicking Submit, you acknowledge that you may be contacted by Fisher Scientific in regards to the feedback you have provided in this form. We will not share your information for any other purposes. All contact information provided shall also be maintained in accordance with our Privacy Policy.